Fort Belvoir, Va. - Consider genetic engineering, molecular biology, nanotechnology, biochemistry and systems biology. What do they all have in common? When you combine the application-orientated sides of these diverse scientific disciplines and throw in a forward engineering approach adopted from mechanical and electronic engineering, you get synthetic biology. This multidisciplined approach enables new medical and environmental diagnostic tools to protect the warfighter.
Synthetic biology is responsible for new classes of inexpensive, rapidly deployable diagnostic devices and is moving towards near real-time surveillance of multiple pathological conditions to protect the warfighter from chemical and biological threats. An article by Defense Threat Reduction Agency-funded researchers, Drs. Shimyn Slomovic, Keith Pardee and Jim Collins, from the Massachusetts Institute of Technology and Harvard University, highlights several milestones and offers insight into where the field is headed. The article, "Synthetic Biology Devices for In Vitro and In Vivo Diagnostics," marks the 100th anniversary of the Proceedings of the National Academy of Sciences and focuses on the evolution of biosensing.
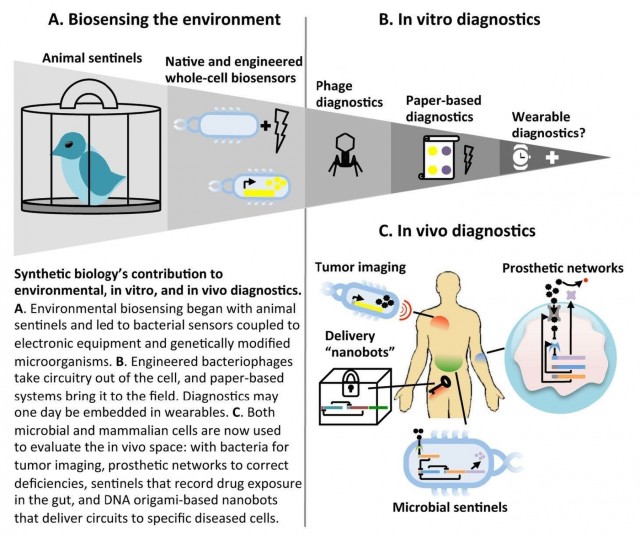
Environmental and whole-cell biosensing
From a historical perspective, biosensing is not a new concept. People have used canaries in coalmines and tracking dogs for centuries to sense environmental hazards. Animal sentinels were scaled down to single living cells in the years leading up to the onset of synthetic biology as a field. The natural biochemical or enzymatic pathways of these first "whole-cell biosensors" were exploited to evaluate the environment.
For example, the photosynthetic activity of electrode-mounted cyanobacteria produced a current that disrupted herbicides in water, alerting to the presence of the chemicals. Such hybrid bio-electrical devices were gradually phased out as scientists introduced bioluminescence-producing gene cassettes into microorganisms, converting them into purpose-built analyte sensors with their own signal output components. However, as microorganisms are progressively modified to include synthetic gene circuitry that allows them to detect arsenic, hydrocarbon pollutants and other compounds, the authors predict a revitalized integration of companion electronic components for rapid field deployment.
The Collin's team also discusses several advances they made as the result of DTRA's Joint Science and Technology funded project managed by Dr. Ilya Elashvili. Notably, they describe how bacterial cells can be programmed with synthetic gene circuitry that transforms them into living RNA-based biosensors, which they reported in the Cell article titled "Toehold Switches: De Novo-Designed Regulators of Gene Expression."
In this paper, the scientists used clever computational algorithms to design RNA "toehold switches" that can be expressed in microorganisms. These RNA-based sensors stem from conventional translational riboregulators, where a toehold hairpin structure complementary to the target is engineered at the upstream of reporter protein mRNA's ribosomal binding site (RBS). The hairpin blocks the RBS binding to ribosome, thereby preventing the initiation of reporter protein synthesis. Upon encountering the complementary target ("trigger") RNA sequence, the hairpin is unzipped (through Watson-Crick base pairing) and the mRNA binding to the RBS is unblocked.
As a consequence, the target RNA sensing is translated to yield the production of fluorescent or colorimetric reporter proteins. The rational de novo, designed from scratch, design of these molecular switches shows great potential for intracellular diagnostics related to disease-specific RNAs. The article proceeds to describe how biosensing machinery has been taken outside of the cell and is currently fulfilling a range of in vitro diagnostic applications.
In vitro diagnostics
Engineered bacteriophages are trailblazing the in vitro diagnostics space. With their natural, strain-specific sensing capabilities, bacteriophages equipped with reporter genes can be deployed to detect, infect and transform pathogens into self-informants that alert of their own presence. These bacterial viruses are inexpensive to make, can be optimized through directed evolution for enhanced target specificity, and are currently in use as pathogen sensors both in clinical settings and in the food industry. Currently the Centers for Disease Control and Prevention uses an engineered plague diagnostic phage, ϕA1122, to detect Yersinia pestis (the etiologic agent of the bubonic plague) in clinical samples in just two hours.
While engineered bacteriophages have begun to take synthetic biology out of the lab, paper-based diagnostics force synthetic biology out of the cell. Cell-free expression systems are commercially available and scientists have also streamlined the production of home-brewed lysates, making it cheaper to translate and screen proteins in vitro. The authors discuss how the same JSTO-funded effort accelerated paper-based systems by freeze-drying coupled transcription/translation reactions onto paper.
In this regard, they cite their accomplishments published in the Cell article, "Paper-Based Synthetic Gene Networks." Here, the authors demonstrate that synthetic gene circuits encoding diagnostic RNA toehold sensors can be freeze-dried in the cocktail as well, and they remain stable for months on end and poised for action until hydrated with a clinical sample or water. For example, this technology allowed Collin's team to construct 24 different Ebola sensors in less than 12 hours with the DNA input costing only 21 per sensor.
The Collins group describes how this technique was used for sensing both small (e.g., glucose) and large molecules (e.g., Ebola virus RNA), including strain-specific Ebola virus sensors. The authors posit that paper-based systems have opened the door to a robust, cheap, mobile and safe means to conduct diagnostic biosensing in low-resource conditions. This new ability has wide applicability to both deployed warfighters and to resource-poor countries.
In vivo diagnostics
Synthetic biology initially aspired to reproduce simple mechanical switches in genetic form, drawing on design principles from mechanical and electrical engineering. Early version genetic toggle switches later grew into increasingly complex circuits that could compute Boolean logic and even facilitate genetic memory. With these advanced circuit design and implementation techniques, researchers realized the potential in using engineered microorganisms to sense the "innerspace" of the human body for real-time diagnostic monitoring.
Collins and his team provide an interesting example of sentinel E. coli engineered with sophisticated toggle--switch circuitry that allows them to patrol the mouse gut, record drug exposure, and report their findings in stool samples. Genetically modified bacteria have also been applied to home in on cancerous growth and relay spatial information via bioluminescence. In the future, such sentinel microorganisms could be critical for early diagnosis and tracking of metastasis, drastically improving the success rates of targeted treatments.
Mammalian cells are also beginning to play important diagnostic and therapeutic roles, carrying out actions that bolster natural biological processes and fix deficient ones. One example given for diagnostic mammalian biosensors describes human cells that were engineered to express the bacterial luxCDABE gene cassette, which allows them to produce a bioluminescent signal following their subcutaneous injection.
The authors also describe the groundbreaking cell-profiler system, in which human cells are equipped with synthetic circuits that allow them to assess the levels of microRNAs that mark specific cancer cells. These sentinels can pick cancer cells out of a "line-up" and either make them glow red or force them to self-destruct by evoking apoptosis. In perhaps the most exquisite example of the therapeutic potential of synthetic biology in mammalian cells, the article discusses a "prosthetic" gene network introduced into encapsulated human cells that are injected into a mouse model to fight gout and tumor lysis syndrome. These cells are able to restore urate homeostasis using genes borrowed from bacteria and fungi, demonstrating a prosthetic gene network concept that holds great potential for future use in humans.
The delivery of genetic constructs into specific cell types and tissues in the human body poses a challenge that is more formidable than drug delivery, as these genes must be expressed once they reach their destinations. Fortunately, researchers are developing an array of delivery means, and the authors detail several of these approaches.
"Nanobots" are origami DNA barrels that can be loaded with genetic content to be delivered to specific cell types whose surface receptor keys open locks found on the bot's hinges. For delivery to tumors, the authors envision the conjugation of nanobots to flagellated commensal bacteria whose self-propulsion and natural tumor-homing attributes could raft the bots to cancerous growth. In addition to adeno-associated viruses and self-assembled virus-like particles, which presently play a prominent role in delivery, biological vesicles have begun to attract much attention.
These lipid pockets can be produced from Gram-negative bacteria and even from exosomes secreted by mammalian cells. Such vesicles are loaded with genetic material and outfitted with cancer-specific antibodies that allow cell-specific delivery and expression of fluorescent reporters, toxins, and anti-cancer small interfering RNAs. The Collins team floats the intriguing prospect of biological vesicles that could be automatically loaded with the appropriate genetic material by "manufacturing" cell lines, which would drastically improve the efficiency and rate of their production.
Synthetic biology has come a long way since its inception, and with it, our ability to engineer novel biological functions, sense the natural environment, diagnose our ailments, and monitor our bodies' internal environment. Biosensing has evolved from animal sentinels to synthetic gene circuits fixed on paper, allowing us to take diagnostics out of the lab and into the field and protect the warfighter. Bacterial, mammalia, and artificial cells are engineered to patrol our insides and inform us of disease-indicating perturbations, and in some cases, to dynamically respond and correct these negative states allowing strong medical counter measures for the warfighter, while having applicability to a wide-range of aliments the world faces.

Social Sharing